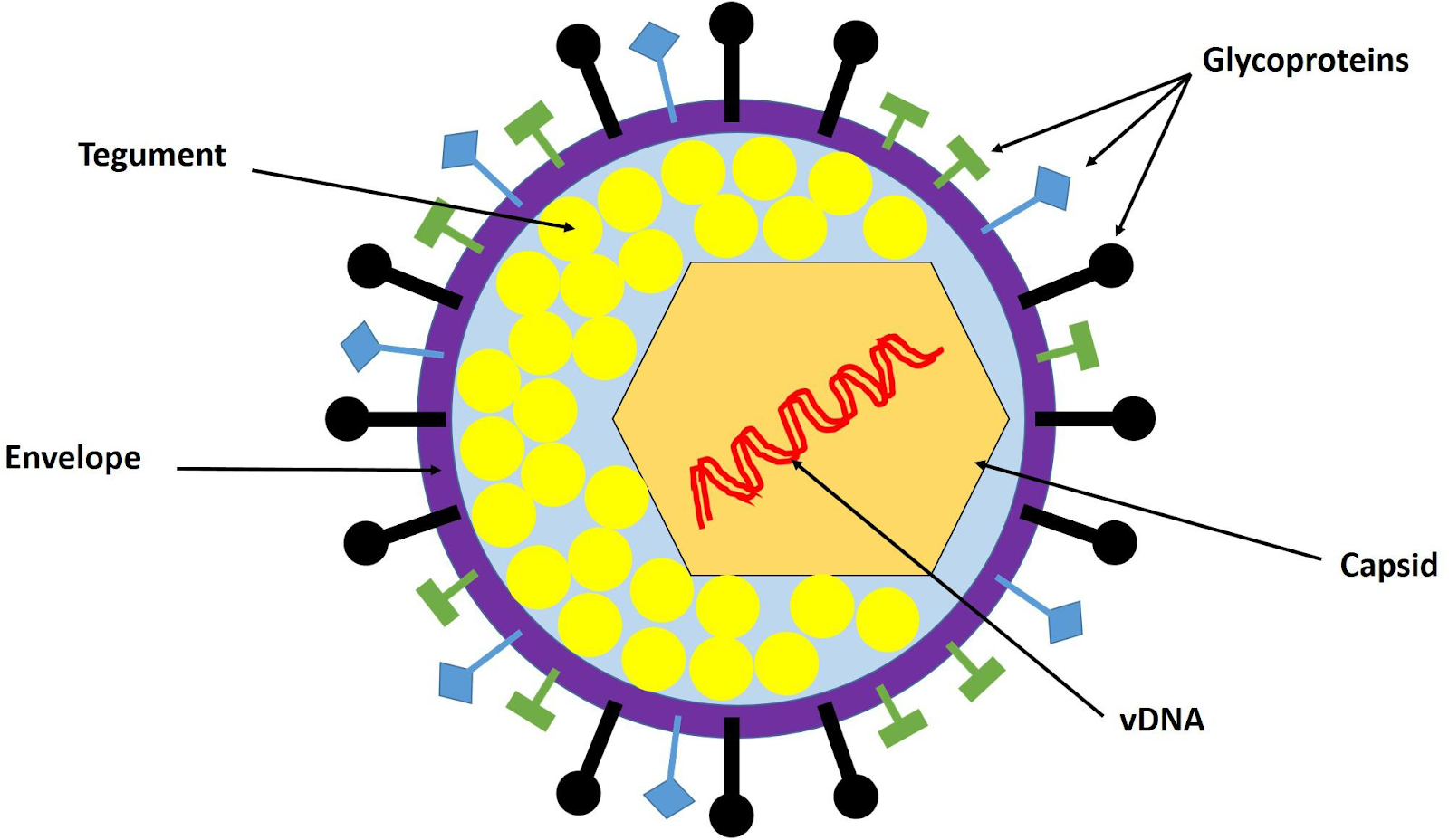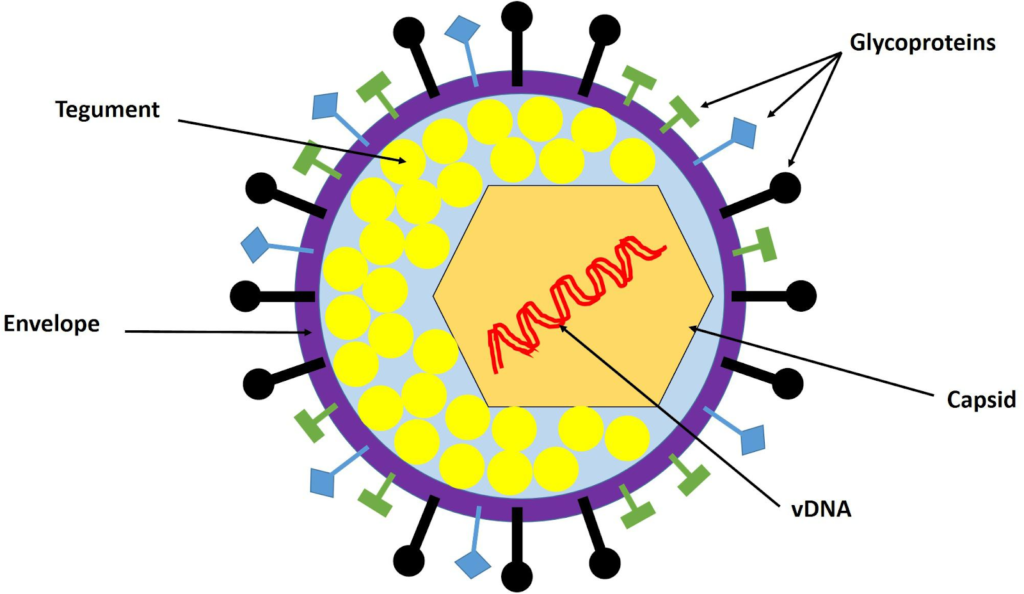
Herpes Simplex Virus (HSV) has a multilayered enveloped structure typical of herpesviruses, with a double-stranded DNA genome packaged inside an icosahedral capsid, surrounded by a tegument and then by a lipid envelope. It features a large, enveloped icosahedral virion about 150 nm to 200 nm in diameter.
Structural Organization of Herpes Simplex Virus
Four Main Structural Layers
From inside to outside, the HSV virion consists of:
1) Core (Genome)
The innermost core contains a large, linear, double-stranded DNA genome (150 to 152 kbp long), tightly packaged as a left-handed spool within the capsid, arranged in a toroidal shape.
2) Capsid
Surrounding the core is an icosahedral capsid (~100-125 nm diameter) made of 162 capsomeres (150 hexons and 12 pentons), forming the nucleocapsid that protects the genome.
3) Tegument Layer
An amorphous protein tegument lies between the capsid and envelop, containing proteins like VP16 (for replication) and VHS (for host shutoff).
4) Envelope Features
The outermost lipid bilayer envelope, derived from host cell membranes, is studded with glycoproteins (e.g., spikes ~8 nm long) that enable cell attachment and entry.
Overall Particle Shape and Size
HSV virions are roughly spherical or pleomorphic, with a total diameter of about 150 200 nm in electron microscopy.
The final structure is a large, enveloped DNA virus with a “core-capsid-tegument-envelope” organization, giving the characteristic “spiky ball” appearance under EM.
Differences in Structure Between HSV-1 and HSV-2
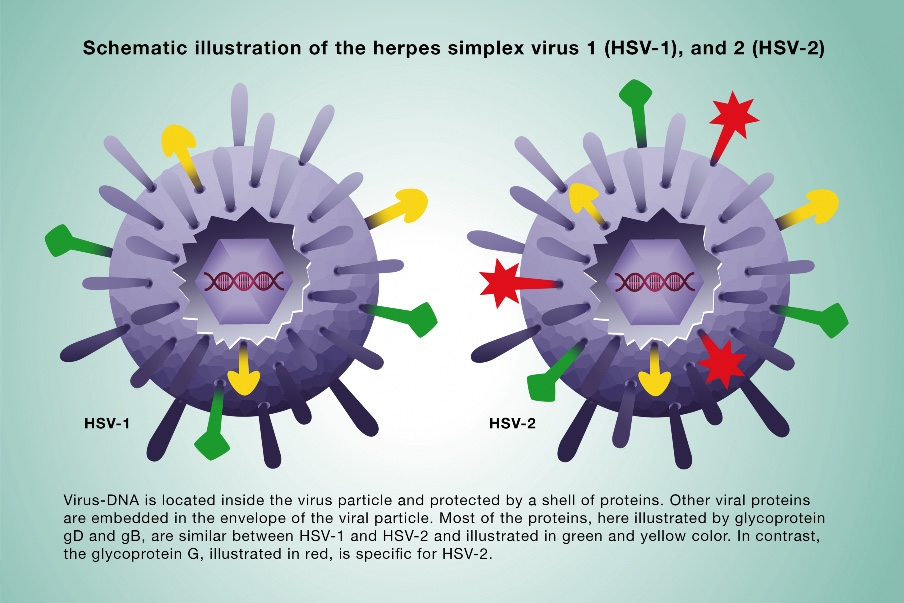
HSV-1 and HSV-2 share nearly identical overall structures as enveloped, icosahedral DNA viruses, but subtle differences exist mainly in glycoproteins and genome sequences that influence cell binding and tropism.
Overall Similarity
Both feature a double-stranded DNA genome in a toroidal core, an icosahedral capsid (150 hexons, 12 pentons), tegument proteins, and a glycoprotein-studded lipid envelope (~180 nm diameter).
Key Differences
Glycoproteins: HSV-1 relies more on 6-0 and 2-3-0 sulfations in heparan sulfate for attachment via gC and gB; HSV-2 has lower dependence but stronger overall HS binding.
Genome: ~80 proteins encoded; HSV-1 and HSV-2 show 50% to 70% homology, with variations in non-essential genes affecting neurovirulence and recurrence.
Structural Impact
These glycoprotein variances lead to HSV-1 favoring oral sites (trigeminal ganglia) and HSV-2 genital sites (sacral ganglia), despite morphological similarity under electron microscopy.
What Glycoproteins Differ Between HSV-1 and HSV-2?
HSV-1 and HSV-2 glycoproteins are highly similar overall, with both viruses encoding about 12 glycoproteins involved in attachment, entry, and spread, but key differences in sequence, structure, and function distinguish them.
Glycoprotein C (gC)
gC-1 (HSV-1) binds heparan sulfate (HS) more strongly via positively charged residues near a Cys127-Cys144 loop, aiding initial attachment; gC-2 (HSV-2) shows weaker HS binding and less reliance on gC for adsorption, shifting more role to gB.
Glycoprotein D (gD)
gD N-termini differ at seven amino acid positions, enhancing HSV-2 gD’s fusion activity with nectin-2 and endogenous CHO receptors compared to HSV-1 gD, which prefers HVEM and nectin-1.
Glycoprotein B (gB)
gB contributes more to HSV-2 attachment due to gC’s reduced role; its cytoplasmic tail in HSV-2 inhibits fusion more than in HSV-1 under certain receptors.
Glycoprotein G (gG)
gG is type-specific; it is absent or non-immunogenic in HSV-1 but immunogenic in HSV-2, enabling serological distinction.
Other Glycoproteins
gB, gH, and gL ratios are similar, but overall glycoprotein diversity is lower in HSV-2 strains; gA/gB shows electrophoretic mobility shifts based on host cells.