Structure and Life Cycle Of Human Papillomavirus (HPV)
Human papillomavirus (HPV) has a small, non-enveloped, icosahedral DNA structure from the Papillomaviridae family. It measures 52-55 nm in diameter and consists of a protein capsid surrounding a circular, double-stranded DNA genome of about 7500 to 8000 base pairs. It primarily infects epithelial cells in skin and mucous membranes. Here’s a breakdown of its structure and life cycle.
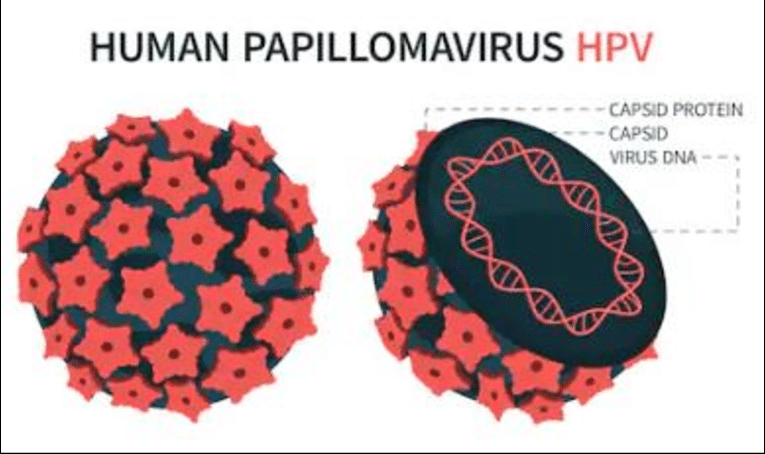
Viral Structure
Capsid Composition
The capsid contains 72 pentameric capsomers made primarily from the L-1 protein (55 kDa, about 80% of viral protein), with fewer L-2 proteins (70 kDa) embedded inside. L1 forms the outer shell for host cell attachment, while L2 aids genome packaging during assembly.
Genome Organization
The DNA is complexed with cellular histones and includes eight open reading frames (ORFs) divided into early (E1-E7 for replication), late (L1-L2 for structure), and a non-coding long control region (LCR) for regulation. This setup enables the virus to hijack host machinery for replication.
Role of L1 and L2 Proteins in HPV Capsid Assembly
L1 and L2 proteins are essential for HPV capsid assembly, with L1 forming the primary structure and L2 enhancing stability and function. L1 self-assembles into virus-like particles, while L2 integrates at substoichiometric levels to support infectious processes.
L1 Protein Role
The major capsid protein L1 comprises 360 copies arranged as 72 pentamers on a T=7 icosahedral lattice, forming the outer shell of the 55-nm virion. L1 alone can spontaneously assemble into empty virus-like particles (VLPs) that mimic native capsids, enabling host cell attachment via surface epitopes. Its pentameric structure provides the scaffold for the entire capsid architecture.
L2 Protein Role
L2, the minor capsid protein, binds to L1 pentamers in a 5:1 stoichiometry via a specific C-terminal domain (amino acids ~396-439 in HPV11), involving hydrophobic interactions resistant to salt and mild denaturants. L2 does not form VPLs independently but co-assembles with L1 to promote ordered T=1 or T=7 particle formation, particularly under acidic conditions (pH 5.2 to 5.4). It embeds internally, aiding DNA packaging and genome delivery during infection.
Co-Assembly Dynamics
Critical residues at the L1-L2 interface ensure precise incorporation; mutations here reduce L2 levels in capsids and lower infectivity. L2 enhances L1 assembly efficiency and stabilizes the genome within the capsid for endosomal trafficking. This interplay is conserved across HPV types, making L2 a target for broad antivirals.
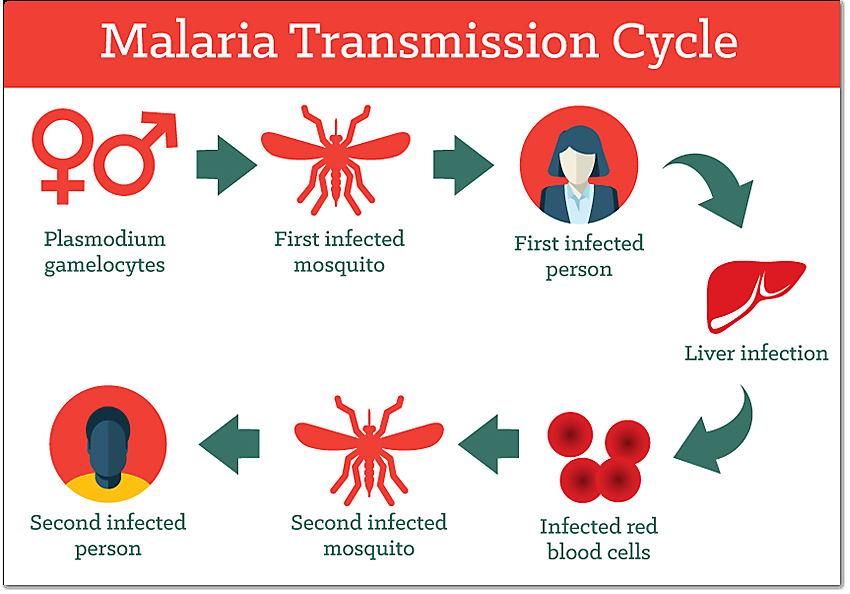
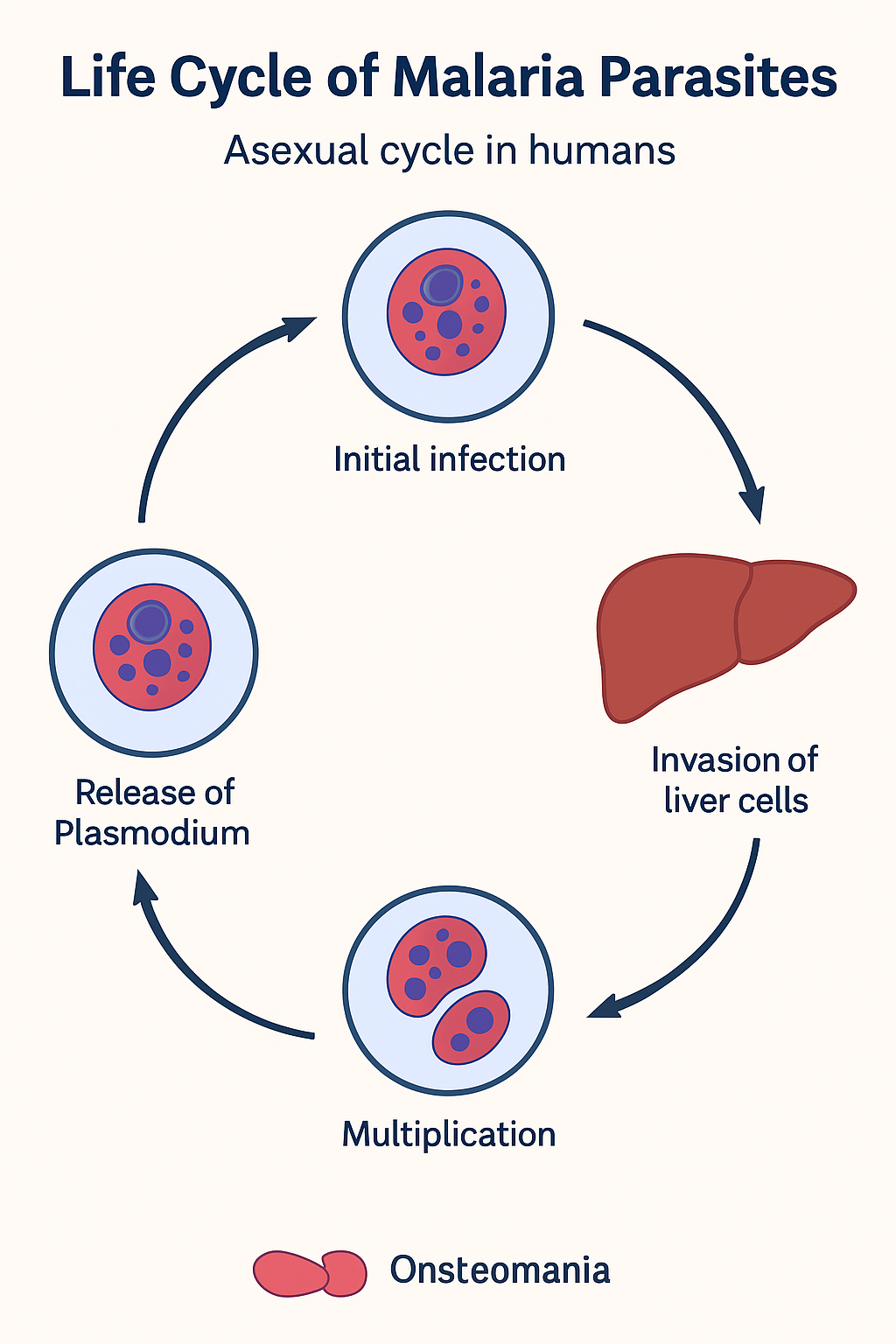
Life Cycle of Human Papillomavirus
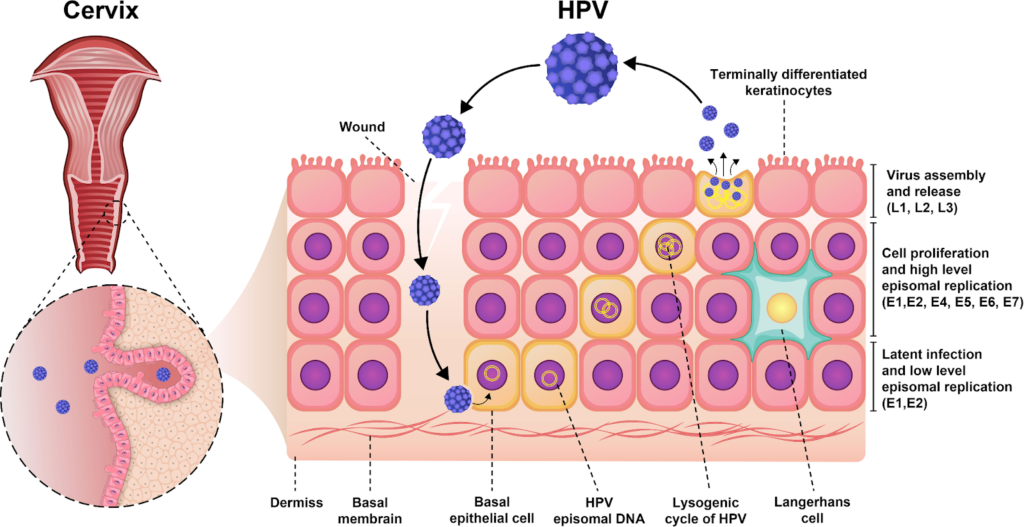
HPV’s life cycle is tightly linked to the differentiation of epithelial keratinocytes, spanning entry in basal cells to virion release from surface layers. It relies entirely on host cell machinery without inducing overt cell lysis, enabling persistent infection.
Basal Layer Entry
Virus enters basal keratinocytes via microtrauma, using heparan sulfate for attachment and L1/L2 proteins for endocytosis into endosomes. The genome escapes as an episome (50-100 copies/cell) via L2-mediated trafficking, establishing low-level maintenance replication driven by E1/E2 proteins synchronized with host S-phase.
Genome Amplification
Infected basal daughters divide; one retains the stem cell niche while others migrate suprabasally and differentiate. E6/E7 oncoproteins reactivate host DNA replication in these post-mitotic cells by degrading p53/pRb, enabling vegetative amplification to thousands of genome copies per cell.
Late Gene Expression
Differentiation cues (e.g., in spinous/granular layers) upregulate late promoters, inducing L1/L2 capsid proteins and E4 for genome release. Progeny virions assemble around histone-complexed DNA, maturing in the nucleus before desquamation sheds infectious particles.
Role of E6 and E7 Proteins in Regulating HPV Life Cycle Stages
HPV E6 and E7 oncoproteins regulate the viral life cycle by overriding host cell cycle controls, enabling genome replication in the differentiating keratinocytes. They ensure episomal maintenance in basal cells and amplify DNA in suprabasal layers by degrading key tumor suppressors.
Episomal Maintenance
In basal keratinocytes, E6 binds E6AP to ubiquitinate and degrade p53, preventing apoptosis and DNA damage responses that could eliminate the low-copy episome (50-100 genomes/cell). E7 binds pRb, freeing E2F transcription factors to drive S-phase entry and support E1/E2-mediated maintenance replication synchronized with cell division.
Vegetative Amplification
As cells differentiate and migrate suprabasally, normal G1/S arrest halts host replication; E6/E7 counteract this by sustaining pRb inactivation and p53 loss, reactivating DNA synthesis for vegetative phase amplification to thousands of genome copies. This creates a pseudomiotic environment permissive for late gene expression (L1/L2) without cell death.
Late Stage Support
Their activities indirectly promote differentiation signals for capsid assembly and genome packaging while preventing premature cell death; mutants lacking functional E6/E7 fail to maintain episomes long-term or amplify DNA. This hijacking links productive infection to epithelial stratification.
Role of E1 and E2 Proteins in HPV DNA Replication
HPV E1 and E2 proteins are primary viral factors driving all phases of DNA replication, from episomal maintenance to vegetative amplification. E1 acts as the origin-binding helicase, while E2 recruits and regulates it at the viral origin.
E1 Functions
E1, an ATP-dependent DNA helicase, recognizes the ATP-rich origin sequence via its C-terminal domain, unwinds DNA at the replication fork, and recruits host replication proteins like MCM and DNA polymerase alpha to initiate bidirectional replication. It forms hexameric rings on single-stranded DNA, traveling 3’->5’ to separate strands during S-phase.
E2 Functions
E2 binds as dimers to four palindromic sites flanking the E1 binding site in the upstream regulatory region (URR), acting catalytically to tether E1 and form the pre-replication complex without a direct helicase loader. It stabilizes E1 at the origin, enhances specificity, inhibits non-specific binding, and modulates replication levels to prevent over-replication.
Replication Phases
Maintenance: Low E1/E2 levels copy the episome (50 to 100 copies/cell) once per basal cell cycle.
Amplification: Upregulated in differentiating cells, E1/E2 drive thousands of genome copies via repeated initiations.
Partitioning: E2 binds host chromosomes to segregate episomes to daughter cells.